Christmas has become the time of year that new computer games are released and this year it is no different. Many people will wake up on Christmas morning to the sound of games such as FIFA, Call of Duty and the many incarnations of dance game available. But is there a way that all this game time could be used to solve scientific problems? Last night I explored 3 online games that do exactly that!
These 3 games are free to play and use the power of crowd-sourcing to solve problems where computer programming has its limits. The best thing about these games is that the solution is not pre-determined. The power of the human mind is used to see patterns and come up with strategies to solve real-life unsolved puzzles.
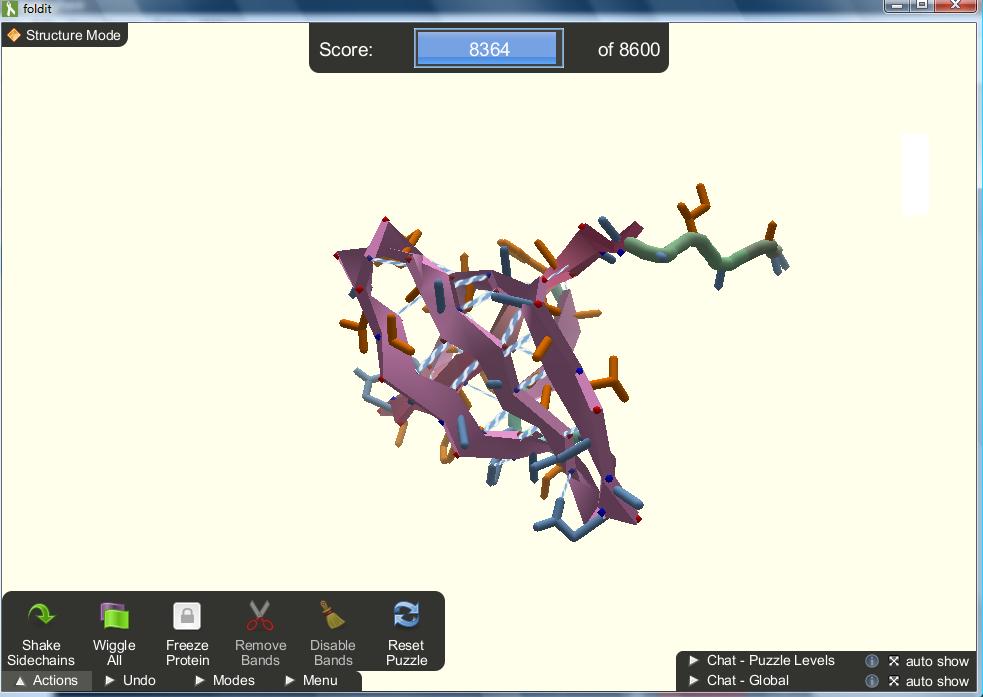
Foldit is the most successful of these games to date with 6 scientific publications and counting. Foldit players are co-authors on these papers so if you want to contribute to a top scientific journal, this is the game for you! Players are given a protein and to gain points they need to determine its structure. Proteins fold into their most stable form but this is hard to determine using computer programs on their own. Proteins which are linked to disease are often targets of drugs. To make drugs that work properly, the correct protein structure needs to be known. Otherwise, the drugs will not interact effectively with the protein.
Players can work individually or in groups and receive points for creating stable protein structures. As part of an experiment in 2010, Foldit players outperformed the structure prediction computer program, Rosetta, on 5 out of 10 puzzles given. A lot of these players had no background in Biochemistry and yet they were able to do beat a state-of-the-art computer program. Since then, Foldit players have solved the structure of a retroviral protease monomer which is a protein made by an AIDS–causing monkey virus, M-PMV. Two monomers of this protein bond together to form an active protein involved in viral maturation and proliferation. Scientists have worked on this problem for over a decade as the monomer is a drug target.
Another aspect of Foldit is that players can create ‘Recipes’ which are algorithms (instructions) they develop. Amazingly one recipe created by the players over a number of months, Blue Fuze, was more effective than an algorithm, Fast Relax, created by scientists over the same period of time. Both algorithms were designed to achieve a more stable protein structure (more points in Foldit game!).

The second game I played last night was eteRNA which is similar to Foldit but you are determining RNA structure. There haven’t been any publications to date resulting from eteRNA but it has been online for a shorter time than Foldit. Players can take part in the Lab once they reach 10,000 points. I joined yesterday and have 2,510 so far so I’ve a bit of a way to go. When you’re good enough to be in the Lab, you compete with other players to figure out how to make stable RNA structures. The best proposals are tested in an actual lab and if they work in vitro, you get bonus points!
I chatted with the number 1 eteRNA player in the world to see what motivated players to spend time on these online games. Eli Fisker accumulated 490,000 points over the past year and said “I started [playing] for fun and science. It is the best game I ever played”. You can’t get a better recommendation than that.

The final game I tried was Phylo which uses brain power to align multiple sequences of DNA, RNA and protein. Again computer programs have some constraints in this areas. Players are given sequences from different species and asked to find the best alignment. These sequences are actual pieces of DNA, RNA or proteins that have been linked to genetic disorders. Phylo state on their website that “Every alignment is received, analyzed, and stored in a database, where it will eventually be re-introduced back into the global alignment as an optimization.”
There are good tutorials included with all of the games so I’d encourage anyone interested to give these a shot. My favourite to play was EteRNA but I was getting the hang of Foldit towards the early hours of last night. Phylo didn’t grab my attention as much but that’s probably down to personal taste.
Overall my journey into the world of crowd-sourcing science games was fulfilling. I enjoyed the games but most of all I didn’t feel like I was wasting my time. There is a lack of the usual guilt associated with online games. It takes a lot of time to get to the standard where you’d be contributing to scientific discovery. Knowing that anyone has the potential to do just that is what makes these games revolutionary.
Happy Christmas from Science Calling!
![]() Cooper, S., Khatib, F., Treuille, A., Barbero, J., Lee, J., Beenen, M., Leaver-Fay, A., Baker, D., Popović, Z., & players, F. (2010). Predicting protein structures with a multiplayer online game Nature, 466 (7307), 756-760 DOI: 10.1038/nature09304
Cooper, S., Khatib, F., Treuille, A., Barbero, J., Lee, J., Beenen, M., Leaver-Fay, A., Baker, D., Popović, Z., & players, F. (2010). Predicting protein structures with a multiplayer online game Nature, 466 (7307), 756-760 DOI: 10.1038/nature09304
Khatib, F., DiMaio, F., Cooper, S., Kazmierczyk, M., Gilski, M., Krzywda, S., Zabranska, H., Pichova, I., Thompson, J., Popović, Z., Jaskolski, M., & Baker, D. (2011). Crystal structure of a monomeric retroviral protease solved by protein folding game players Nature Structural & Molecular Biology, 18 (10), 1175-1177 DOI: 10.1038/nsmb.2119
Khatib, F., Cooper, S., Tyka, M., Xu, K., Makedon, I., Popovic, Z., Baker, D., & Players, F. (2011). From the Cover: Algorithm discovery by protein folding game players Proceedings of the National Academy of Sciences, 108 (47), 18949-18953 DOI: 10.1073/pnas.1115898108
Top image: Arpingstone / Wikimedia Commons

Last night i played Foldit thanks to your article “The New Revolution…”,it was something new to me as kind of game i never played before but it was very fun and interesting.
Thanks
Glad you enjoyed it!